 |
 |
AbstractPurposeThe aim is to study the dependence of deformable based auto-segmentation of head and neck organs-at-risks (OAR) on anatomy matching for a single atlas based system and generate an acceptable set of contours.
MethodsA sample of ten patients in neutral neck position and three atlas sets consisting of ten patients each in different head and neck positions were utilized to generate three scenarios representing poor, average and perfect anatomy matching respectively and auto-segmentation was carried out for each scenario. Brainstem, larynx, mandible, cervical oesophagus, oral cavity, pharyngeal muscles, parotids, spinal cord, and trachea were the structures selected for the study. Automatic and oncologist reference contours were compared using the dice similarity index (DSI), Hausdroff distance and variation in the centre of mass (COM).
ResultsThe mean DSI scores for brainstem was good irrespective of the anatomy matching scenarios. The scores for mandible, oral cavity, larynx, parotids, spinal cord, and trachea were unacceptable with poor matching but improved with enhanced bony matching whereas cervical oesophagus and pharyngeal muscles had less than acceptable scores for even perfect matching scenario. HD value and variation in COM decreased with better matching for all the structures.
ConclusionImproved anatomy matching resulted in better segmentation. At least a similar setup can help generate an acceptable set of automatic contours in systems employing single atlas method. Automatic contours from average matching scenario were acceptable for most structures. Importance should be given to head and neck position during atlas generation for a single atlas based system.
IntroductionRadiotherapy treatment planning is a time consuming process. The target volumes and organs-at-risks (OAR) are manually delineated for treatment plan generation. In head and neck cancers, accurate contour delineation is essential for a good treatment outcome. Manual delineation of contours is a cumbersome task especially in busy departments [1-3]. It is at this stage the role of auto-segmentation in the planning process has become important. The concept of auto-segmentation of OAR and clinical target volumes (CTV) has been introduced for faster delineation of contours and also to reduce inter-observer variation [3,4]. Auto-segmentation algorithms may be atlas based, model based or hybrid based [4]. Predominantly all the systems use atlas based deformable image registration (DIR) for contour generation. DIR represents the transformation between two image sets where the voxels of the moving image set are warped to match the voxels of the target image set and is represented by a deformation vector field. Several applications of DIR are documented in literature [5]. In addition to contour auto-segmentation, it can be used to determine delivered doses and generate cumulative dose-volume histograms [5,6]. The commercially available systems predominantly employ atlas based systems are for auto-segmentation. Teguh et al. [7] studied atlas based auto-segmentation of CTV and OARs for 12 patients and concluded that it offers high throughput but manual editing is essential. Daisne and Blumhofer [8] have auto-segmented OARs and CTV and compared it with manually delineated contours. They have reported significant time saving for OARs than CTV. Stapleford et al. [9] have shown the reduction in inter-observer variability in the auto-segmented contours. Thomson et al. [10] investigated auto-segmentation of five OARs namely parotids, submandibular glands, larynx, pharyngeal muscles, and cochlea and reported that automatic contours were inaccurate except for the parotid and submandibular gland.
Immobilization is unique for each patient in head and neck radiotherapy. Images of the same patient acquired at different point of time during the course of treatment seldom match exactly due to weight loss and head rotation. It is difficult to attain a perfect match between the library and sample patient. Though DIR accounts for the anatomy changes, errors are introduced for large variations and differences in setup. Many studies have pointed the advantage of multi-atlas over single-atlas based delineation [7,11,12]. Multi atlas based systems typically score over single atlas based method by accounting for mismatch in size and anatomic variation between the library and sample patients by generating an average patient from the library data. Time saving has also been reported with multi atlas based system [12]. However, some of the commercially available systems still continue to use single atlas methods for auto-segmentation. Anatomic similarity is essential for a good auto-segmentation in single atlas based systems. Several studies have reported on atlas based auto-segmentation [7-17]. But, none have determined the dependence of auto-segmentation on anatomic similarity between the sample and atlas patients. In this study, we have analyzed the effect of patient setup and position on the outcome of a single atlas based auto-segmentation. We have created three different levels of matching between the atlas and sample patients to determine (1) the relation between anatomy matching and OAR segmentation accuracy for different structures in head and neck patients, (2) the minimum prerequisite matching required to generate an acceptable set of contours.
Methods and Materials1. Smart segmentation moduleThe Smart segmentation module of Eclipse treatment planning system (version 13.6; Varian Medical Systems, Palo Alto, CA, USA) was used to generate automatic contours. The module utilizes a DIR-based single atlas method for auto-segmentation. For a sample patient, the system helps in identifying the suitable expert case from library data by estimating similarity between sample and library image sets. The estimated вҖҳsimilarityвҖҷ is based on the anatomic geometry of image sets and is generated using the intensity matching between image sets. The similarity is represented by a scale ranging from 1 to 5, with 1 denoting a least match and 5 the best. For a given sample case, similarity value is specified beside each of the available library case. The user can select the library case with highest value. Initially, rigid registration is carried between sample patient and selected expert library patient to correct for gross errors. DIR is then performed to account for anatomical variations and the contours are propagated by deforming the expert case contours. The DIR utilizes a modified accelerated demons algorithm proposed by Wang et al. [18] where the deformation is based on the intensity differences between the images.
2. Patient selection and atlas generationPatients without gross nodal metastasis who underwent treatment for head and neck cancer for the first time were selected retrospectively. The patients were immobilized with five clamp head and neck thermoplastic mask (Orfit Industries, Wijnegem, Belgium). The institution protocol is to acquire a plain image followed by a contrast enhanced image set. Contouring and plan generation are done on the plain image set only. The contrast enhanced set is used only to aid target volume delineation. The CT images were acquired in a Siemens Biograph 16 slice PET-CT scanner (Siemens Medical Systems, Concord, CA, USA) with a slice thickness of 3 mm. The contours that were earlier manually delineated for plan generation were reviewed by an expert team of oncologists and taken as reference. Three atlas sets with 10 patients each were created (Table 1). All patients in a given atlas set had similar neck position. The first atlas set (ATLASEXT) consisted of patients in extended neck position (defined by a sternal notch to chin distance in the range of 13вҖ“14 cm), while the second (ATLASN) and third (ATLASP) atlas sets consisted of patients in neutral neck position (sternal notch to chin distance in the range of 8вҖ“9 cm). A 10 patient sample was used to study the output of the auto-segmentation using the three atlas sets. The patients in the test sample and third atlas ATLASP were the same but, consisted of CT images acquired at different times. All patients in the third atlas set had undergone PET-CT as a part of imaging process for accurate gross tumour delineation. PET-CT image acquisition was done with the patients immobilized exactly as during simulation. The ATLASP included the CT component of PET-CT while the test sample included the planning CT images. The expert team also delineated the contours on the CT component of PET-CT.
3. Auto-segmentationThe oncologist delineated contours on the atlas image sets were transferred to the sample image sets using Smart segmentation module. Three anatomy matching scenarios were created using the ten test sample patients and three atlas sets. The first scenario (poor matching) was created by using the set ATLASEXT. Automatic contours were generated for all ten sample patients using the atlas library case with least similarity value of 1. In case of more than one atlas library case with the least similarity to any sample patient, the library case with large size difference was chosen for auto-segmentation. Poor matching scenario had a total mismatch in head and neck position, size and anatomy between the sample and atlas patients. The second scenario (average matching) was created by using the set ATLASN. Automatic contours were generated for all sample patients using the library case having a similarity value of 3. In case of more than one atlas library case with a similarity value of 3, the one with a similar size was chosen for auto-segmentation. Although the setup is similar, differences in anatomy between patient to patient and a mismatch in size would exist in this scenario. The third scenario (perfect matching) was created using ATLASP. There would be a maximum similarity only if both the image sets are similar. It is nearly impossible to generate this using multiple patients and the easiest way to achieve it was to use two different image sets of a patient in same setup, acquired in a short span of time. In perfect matching scenario, auto-segmentation was carried using data from the same patient. The automatic contours generated for all 10 test sample patients using each of the three scenarios were compared against the reference contours.
4. Contour evaluationWe studied the auto-segmentation of brainstem, mandible, larynx, cervical oesophagus, oral cavity, parotids, pharyngeal muscles, spinal cord, and trachea. The head and neck OAR delineation guidelines [19] were followed for manual delineation of structures. For the sake of simplicity, the pharyngeal muscles were contoured as a single structure, while larynx contours included the supraglottic, glottic, and subglottic regions. The oncologist contours in the test sample images were considered as the gold standard to which the automatic contours for the three scenarios were compared. Dice similarity index (DSI), Hausdorff distance (HD), and variation in the centre of mass (COM) were the metrics used for contour analysis [20]. DSI is a geometric volumetric similarity measure used to determine the degree of overlap of two set of contours. The value can range from 0 to 1. A value of вҖҳ1вҖҷ indicates perfect overlap of contours and вҖҳ0вҖҷ indicates null overlap. If вҖҳAвҖҷ and вҖҳBвҖҷ are two contours, then DSI is defined as
The DSI scores were determined and the scores were analyzed for all the three different scenarios. A DSI score range of 0.60 to 0.80 has been reported among physician drawn contours only [3]. In this study, a DSI score greater than or equal to 0.80 was accepted as good matching criteria [13,21]. HD measures the degree of mismatch between two image sets based on contour boundaries. It is defined as the maximum distance between a point in one image set and the corresponding point on another image set. The HD values were determined using Slicer 3D Software. COM is used to determine the absolute position of contours based on the three-dimensional coordinates generated by the planning system. A perfect match is confirmed by the same set of coordinates. It can help in tracking the position of one set of contours with respect to another and also over a period of time. Variation in COM of the automatic contours with respect to reference contours were studied.
5. Statistical analysisAll statistical analyses were carried out in Microsoft Excel (version 2013). Single factor analysis of variance (ANOVA) was used to test the significance of DSI scores. A p-value of less than 0.05 was considered statistically significant and less than 0.001 as highly significant. In addition, a post-hoc test using Bonferroni approach was carried out if the ANOVA test returned a significant difference. It uses a two sample t-test to ascertain exactly which of the scenarios were different by using three combinations namely scenario 1 versus 2, scenario 1 versus 3, and scenario 2 versus 3.
Results1. DSI scorePerfect matching and poor matching scenarios produced the best and worst results, respectively, for all studied structures. In contrast to other two scenarios, average matching scenario produced mixed results. The DSI scores for all the three anatomy matching scenarios are represented in Fig. 1. Amongst all structures, brainstem had the best DSI score irrespective of anatomy matching and cervical oesophagus had the least DSI score in all the three scenarios. In case of poor match scenario, brainstem contours were acceptable while mandible and oral cavity contours were close to acceptance criteria. Average matching scenario yielded good results for five of the nine structures studied. Scores of cervical oesophagus, larynx, pharyngeal muscle and trachea were below the acceptance threshold. Substantial improvement in automatic contours of mandible, oral cavity, spinal cord, and parotid contours with better anatomy matching were notable. Larynx and trachea were close to threshold while cervical oesophagus and pharyngeal muscles were well below the acceptable criteria. In perfect match scenario, all but cervical oesophagus and pharyngeal muscle had DSI scores greater than 0.80. Overall, automatic delineation of all the structures improved with superior anatomy matching. Fig. 2 depicts the reference contours and auto-segmented contours on sample patients for the different matching scenarios, respectively.
2. Hausdorff distanceThe HD values for all three scenarios are shown in Fig. 3. Brainstem, spinal cord, and trachea were the only structures to have low HD values in poor and average matching scenarios. High HD values for oesophagus, larynx, and pharyngeal muscles were also observed. There were minor reduction in HD values for average matching, while significant reductions were noticed for perfect matching as compared to poor matching scenario. Perfect matching yielded the least HD scores.
3. COM variation
Fig. 4 depicts the absolute variation in the COM for the three matching scenarios. In case of poor matching scenario, all structures except brainstem had significant deviation along the longitudinal direction while parotids and trachea had large shifts in all directions. Cervical oesophagus was the only structure that had large deviation along lateral direction. In average matching scenario, cervical oesophagus, and pharyngeal muscles had considerable shift in COM. Overall shift in COM reduced with improved anatomy matching for all structures. Structures with high DSI scores correspondingly had less deviation in COM. The comparison with the results obtained from few other similar studies in literature are shown in Table 2.
4. Statistical analysisBased on ANOVA test, the three scenarios were statistically different with p-value less than 0.001 for all the structures studied. As the ANOVA test returned a significant difference, a post-hoc test using Bonferroni approach was carried out. The p-values obtained from the ANOVA test and the post-hoc test is depicted in Table 3.
Discussion and ConclusionWe have studied deformable registration based auto-segmentation for different levels of anatomy matching between sample patients and atlas patients using DSI scores, HD, and variation in COM. The effect of patient position on the output of auto-segmentation has not been studied before. Amongst the structures studied, brainstem segmentation was the best with the mean DSI scores greater than 0.80 and low COM variation for all three scenarios. Brainstem auto-segmentation was found to be less dependent on head position and our results were comparable with that of multi atlas systems [8,13,14,16]. Head and neck auto-segmentation challenge have also produced good results for brainstem with four out of five teams achieving a DSI score greater than 0.80 [13]. Daisne and Blumhofer [8] have shown a DSI score of 0.80 but for corrected auto-segmented brainstem contours. Intensity differences existing between brainstem and surrounding brain had resulted in better segmentation. The over estimation along the superior direction and missing parts of brainstem along the inferior direction were reflected in the HD value and shift in COM along longitudinal axis.
Being a high intensity structure surrounded by low intensity structures, mandible could be easily segmented. Studies have reported DSI values in the range of 0.78вҖ“0.98 [13,14,16]. Tsuji et al. [15] have shown that structures like mandible with clearly defined borders in CT had superior auto-segmentation accuracy. However in our case, a less than acceptable mean DSI score was obtained in the poor matching scenario because of huge differences in chin position between the sample and atlas patients and the inability of the system to handle very large deformation accurately. The system over estimated mandible in all scenarios by including teeth but the extrusion reduced with improved bony matching. This resulted in a high HD and COM variation for mandible in poor bony matching scenario. It is essential to have at least a similar head position between atlas and sample patients if not a perfect chin match, for an acceptable mandible contour. Accuracy of oral cavity automatic contours were highly influenced by mandible or head position. Large differences in mandible position as in poor matching scenario resulted in a less than acceptable mean DSI score for oral cavity. High HD values in poor and average matching scenarios were due to the overestimation of automatic contours in posterior and inferior direction. High density structures limited the extrusion of oral cavity contours along the superior and anterior direction. Parotid was another structure that required a similar head position for a good segmentation. The segmentation was not accurate in case of large differences in head position. Our parotid segmentation results were comparable with multi atlas studies which have shown DSI values ranging from 0.71 to 0.89 [9,11,14,16]. The inner lobes of the parotids were partly missed in automatic contours resulting in high HD values in poor and average matching scenario.
Larynx, spinal cord, and trachea segmentation were highly dependent on the neck position. Better the neck similarity more acceptable the automatic contours. This has also been shown by Barley et al. [17] who have reported that structures close to bony anatomy like larynx were highly dependent on the range of available atlas cases. In our case, the system could not exactly deform the structures in patient samples with contrasting neck position to that of atlas patients and also overestimated larynx and trachea at the boundaries along the superior-inferior direction. The relative shift in larynx contours along the longitudinal axis in poor match scenario resulted in high HD value and large shift in COM. The changes in neck position affecting spinal cord auto-segmentation has also been reported by Tsuji et al. [15]. In case of spinal cord, the contours were satisfactory only in average and perfect match scenarios. Being encompassed by bony anatomy resulted in the least HD value in all three scenarios. In poor matching scenario, the variation in COM along vertical axes were high due to inappropriate neck matching while lateral shifts were restricted due to surrounding vertebrae. Although trachea could be easily segmented due to surrounding intensity differences, automatic contours missed parts of it inferiorly and did not include tracheal cartilage in poor and average matching scenarios. Cervical oesophagus and pharyngeal muscle are relatively low intensity structures in a low contrast region and also subject to movement. Hence segmentation becomes difficult and thus had the least DSI score and high HD value among all structures. This might be due to the inherent limitation of intensity based deformable registration algorithms in low contrast regions. Pharyngeal muscle segmentation in poor and average matching scenarios were bad because of the extrusion of automatic contours into pharyngeal cavity. It has been shown that statistical differences were not significant between multi atlas and single atlas methods [11]. However in our case, differences were highly significant between the three scenarios.
An optimal extent of anatomy matching required for a good auto-segmentation has not been defined in literature. In our case, perfect match scenario produced the best results. However, it is practically difficult to create such a scenario in a clinical environment. Based on metrics analysed, results from the average matching scenario could be comparable with that of multi atlas based systems for five of the nine structures studied [8,13,14,16,22]. Larynx and trachea may also be considered as their scores were close to acceptance criteria in the average match scenario. Only cervical oesophagus and pharyngeal muscle contours could not be accepted. Although studies have shown the superiority of multiple atlas based over single atlas based methods, comparable automatic contours for most of the studied structures can be generated by having at least an average matching scenario representing a similar setup as shown in our study. Verification and correction of automatic contours are necessary before plan generation as even studies with multi atlas based systems and model based systems have concluded that automatic contours still required manual intervention [7,8,11,12]. DIR is considered as the universal choice for auto-segmentation as it can account for anatomical variations but accurate deformation is difficult in case of large variations. In addition, deformable errors may also be introduced due to lack of uniform CT image acquisition parameters. Acceptable auto-segmented contours can be obtained with improved matching by customizing the atlases based on head and neck position. From this study we have shown that good automatic contours for all structures except cervical oesophagus and pharyngeal muscle could be obtained if there was at least a similar setup between the sample and atlas patients. The limitations of the study are we have used a small sample size of 10 patients and have not studied time savings by editing the automatic contours.
In conclusion, from the study we have shown that in case of single atlas method, the extent of anatomy matching between the sample and atlas patients plays a decisive role in the autosegmentation process. At least a similar setup is an essential pre-requisite to generate an acceptable set of automatic contours in single atlas based systems. While generating an atlas importance should also be given to the head and neck position.
Fig.В 1.The mean dice similarity index (DSI) scores with error bars for structures from the three matching scenarios. 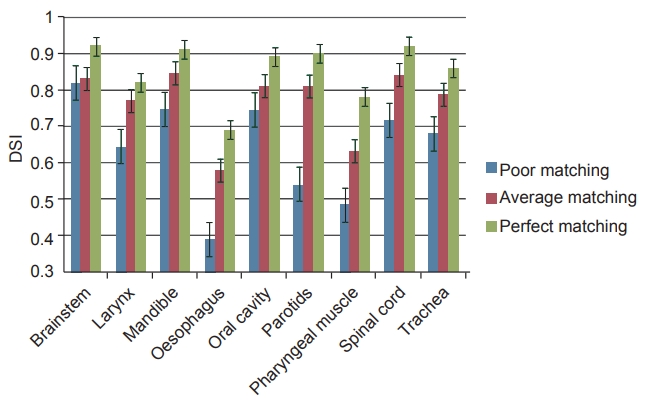
Fig.В 2.Axial and sagittal views of reference and auto contours overlaid on a sample patient for the three scenarios. 
Fig.В 3.The mean Hausdorff distance (HD) values with error bars for structures from the three matching scenarios. 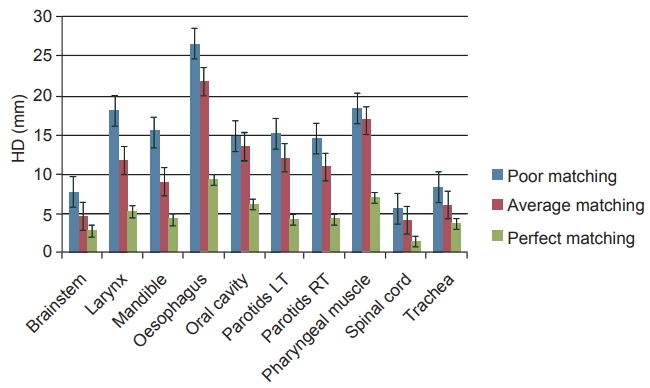
Fig.В 4.Centre of mass for three scenarios: S1, S2, and S3 represent the poor, average, and perfect matching scenarios, respectively. 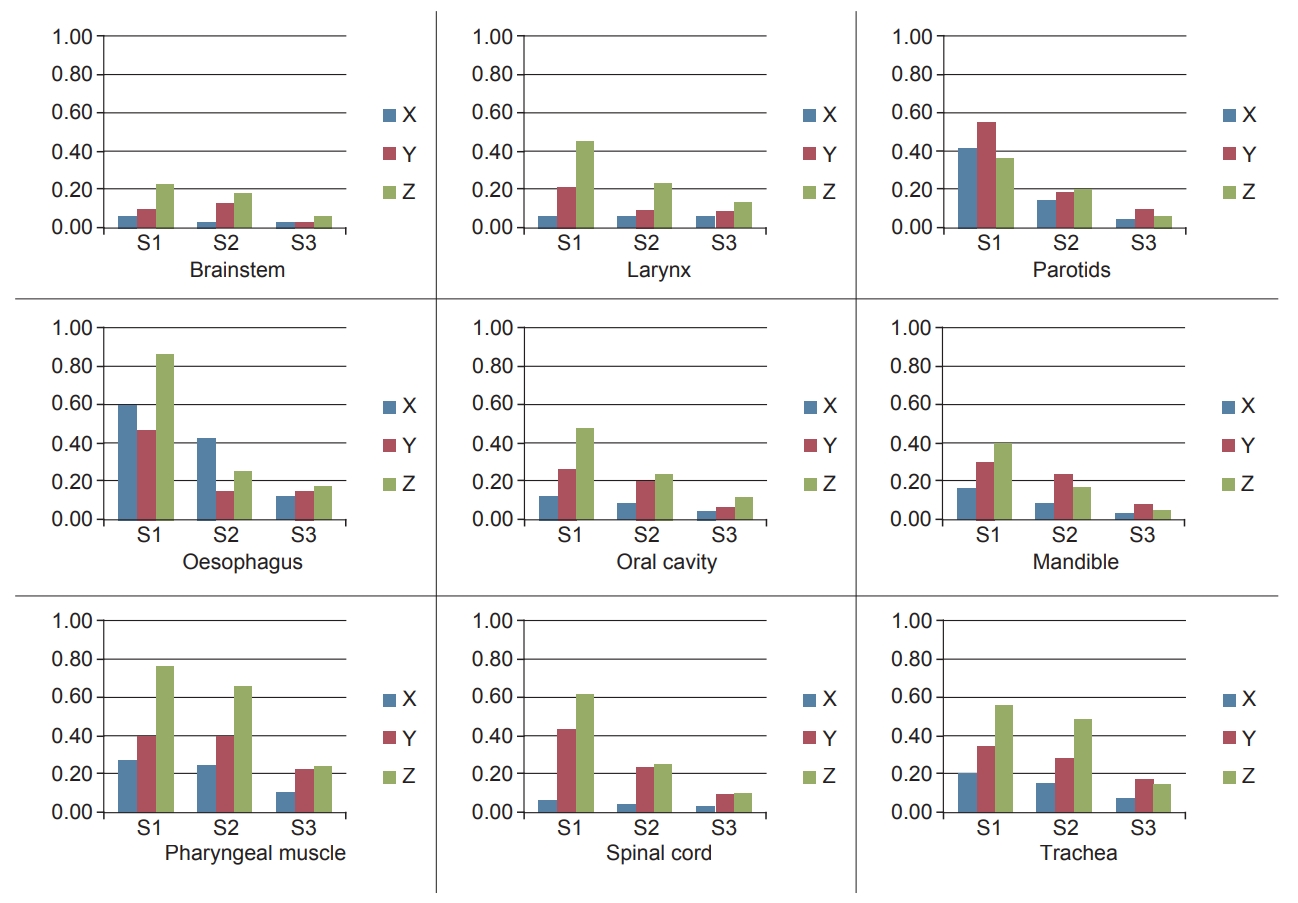
TableВ 1.Patient characteristics of the atlas and sample patients
TableВ 2.Comparison between the results from our study and other similar studies in literature
TableВ 3.The obtained p-values for ANOVA and post-hoc tests References1. Chao KS, Bhide S, Chen H, et al. Reduce in variation and improve efficiency of target volume delineation by a computer-assisted system using a deformable image registration approach. Int J Radiat Oncol Biol Phys 2007;68:1512вҖ“21.
2. Reed VK, Woodward WA, Zhang L, et al. Automatic segmentation of whole breast using atlas approach and deformable image registration. Int J Radiat Oncol Biol Phys 2009;73:1493вҖ“500.
3. Sharp G, Fritscher KD, Pekar V, et al. Vision 20/20: perspectives on automated image segmentation for radiotherapy. Med Phys 2014;41:050902.
4. Lim JY, Leech M. Use of auto-segmentation in the delineation of target volumes and organs at risk in head and neck. Acta Oncol 2016;55:799вҖ“806.
5. Ayyalusamy A, Vellaiyan S, Shanmugam S, et al. Feasibility of offline head & neck adaptive radiotherapy using deformed planning CT electron density mapping on weekly cone beam computed tomography. Br J Radiol 2017;90:20160420.
6. Kadoya N. Use of deformable image registration for radiotherapy applications. J Radiol Radiat Ther 2014;2:1042.
7. Teguh DN, Levendag PC, Voet PW, et al. Clinical validation of atlas-based auto-segmentation of multiple target volumes and normal tissue (swallowing/mastication) structures in the head and neck. Int J Radiat Oncol Biol Phys 2011;81:950вҖ“7.
8. Daisne JF, Blumhofer A. Atlas-based automatic segmentation of head and neck organs at risk and nodal target volumes: a clinical validation. Radiat Oncol 2013;8:154.
9. Stapleford LJ, Lawson JD, Perkins C, et al. Evaluation of automatic atlas-based lymph node segmentation for head-and-neck cancer. Int J Radiat Oncol Biol Phys 2010;77:959вҖ“66.
10. Thomson D, Boylan C, Liptrot T, et al. Evaluation of an automatic segmentation algorithm for definition of head and neck organs at risk. Radiat Oncol 2014;9:173.
11. Yang J, Beadle BM, Garden AS, et al. Auto-segmentation of low-risk clinical target volume for head and neck radiation therapy. Pract Radiat Oncol 2014;4:e31вҖ“7.
12. Sjoberg C, Lundmark M, Granberg C, Johansson S, Ahnesjo A, Montelius A. Clinical evaluation of multi-atlas based segmentation of lymph node regions in head and neck and prostate cancer patients. Radiat Oncol 2013;8:229.
13. Raudaschl PF, Zaffino P, Sharp GC, et al. Evaluation of segmentation methods on head and neck CT: autosegmentation challenge 2015. Med Phys 2017;44:2020вҖ“36.
14. Han X, Hoogeman MS, Levendag PC, et al. Atlas-based autosegmentation of head and neck CT images. In : Metaxas D, Axel L, Fichtinger G, Szekely G, editors. Medical image computing and computer-assisted intervention. Heidelberg: Springer; 2008, p. 434вҖ“41.
15. Tsuji SY, Hwang A, Weinberg V, Yom SS, Quivey JM, Xia P. Dosimetric evaluation of automatic segmentation for adaptive IMRT for head-and-neck cancer. Int J Radiat Oncol Biol Phys 2010;77:707вҖ“14.
16. Walker GV, Awan M, Tao R, et al. Prospective randomized double-blind study of atlas-based organ-at-risk autosegmentation-assisted radiation planning in head and neck cancer. Radiother Oncol 2014;112:321вҖ“5.
17. Barley S, Antoine C, Webster G, et al. Atlas-based autocontouringвҖ“balancing accuracy with efficiency in OnQ rts. Eur Oncol Haematol 2014;10:98вҖ“101.
18. Wang H, Dong L, O'Daniel J, et al. Validation of an accelerated 'demons' algorithm for deformable image registration in radiation therapy. Phys Med Biol 2005;50:2887вҖ“905.
19. Brouwer CL, Steenbakkers RJ, Bourhis J, et al. CT-based delineation of organs at risk in the head and neck region: DAHANCA, EORTC, GORTEC, HKNPCSG, NCIC CTG, NCRI, NRG Oncology and TROG consensus guidelines. Radiother Oncol 2015;117:83вҖ“90.
20. Jameson MG, Holloway LC, Vial PJ, Vinod SK, Metcalfe PE. A review of methods of analysis in contouring studies for radiation oncology. J Med Imaging Radiat Oncol 2010;54:401вҖ“10.
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||

 |
 |