Harnessing genome-wide association studies to minimize adverse radiation-induced side effects
Article information
Abstract
Radiotherapy is used as definitive treatment in approximately two-thirds of all cancers. However, like any treatment, radiation has significant acute and long-term side effects including secondary malignancies. Even when similar radiation parameters are used, 5%–10% of patients will experience adverse radiation side effects. Genomic susceptibility is thought to be responsible for approximately 40% of the clinical variability observed. In the era of precision medicine, the link between genetic susceptibility and radiation-induced side effects is further strengthening. Genome-wide association studies (GWAS) have begun to identify single-nucleotide polymorphisms (SNPs) attributed to overall and tissue-specific toxicity following radiation for treatment of breast cancer, prostate cancer, and other cancers. Here, we review the use of GWAS in identifying polymorphisms that are predictive of acute and long-term radiation-induced side effects with a focus on chest, pelvic, and head-and-neck irradiation. Integration of GWAS with “omic” data, patient characteristics, and clinical correlates into predictive models could decrease radiation-induced side effects while increasing therapeutic efficacy.
Introduction
Radiation therapy (also called radiotherapy) is a localized cancer treatment that targets cancer cells with high doses of radiation. More than half of patients with cancer, including children with cancer, will be treated with radiotherapy [1]. Acute radiation-induced side effects include dermatitis and fatigue, while long-term side effects include increased risk for secondary malignancies. Site-specific acute side effects such as mucositis, esophagitis, and enteritis, may develop when treating head and neck cancers, lung cancers, and gastrointestinal (GI) cancers, respectively. Patients treated for breast cancer and lung cancer may also develop irreversible cardiotoxicity and pulmonary fibrosis after thoracic irradiation [2,3]. Normal tissue toxicity determines the maximum radiation dose delivered to tumors. To increase the therapeutic ratio of radiation, radiation oncologist and their teams individualize patient’s treatment plans by selecting the appropriate treatment modality, radiation dose, fractionation, and target volumes. Recent phase II–III clinical trials have focused on comparing treatment modalities such as ultra-hypofractionation versus conventionally fractionated therapy, and reducing effective doses to decrease acute and long-term side effects [4,5]. Despite this customization, 10%–20% of patients receiving radiotherapy develop grade 2 toxicity and 2%–5% develop grade 3 toxicity [6,7]. Genetic variability, in part, explains this clinical variability.
It is estimated that genetic variability may account for approximately 40% of the variability seen in treatment side effects, while lifestyle risk factors, not limited to obesity, smoking and drinking, account approximately for 60% of the variability [8–10]. Single nucleotide polymorphisms (SNPs), gene deletions, epigenetic differences and differential gene expression are markers of genetic variability. Genome-wide association studies (GWAS) use an agnostic approach to identify SNPs associated with a predetermined outcome such as normal tissue toxicity. Though the majority of GWAS have focused on polymorphisms that increase the risk of developing primary cancers and tumor radiosensitivity, there is evidence that polymorphisms that increase the risk of radiation-induced cancers differ from those that increase the risk of primary cancers [11,12]. Therefore, genetic markers of normal tissue toxicity secondary to radiation are needed to further personalize treatments and improve outcomes.
Historically, genes that increase normal tissue toxicity following radiation were discovered through studies of rare monogenetic diseases such as Li-Fraumeni syndrome, retinoblastoma, neurofibromatosis type 1, ataxia telangiectasia, and Fanconi’s anemia. These diseases not only raised the knowledge about genetic predisposition of normal tissue toxicity, but also provided a biological framework for future discoveries, such as those informed by candidate gene approaches [13]. Candidate gene approaches further identified polymorphisms associated with genes involved in DNA repair, oxidation, and cytokine synthesis that increase the risk for radiation-induced side effects [7]. The development of SNP microarrays paved the way for non-priori identification of additional polymorphisms through GWAS approaches. To increase the power that is needed to identify polygenic causes of radiotoxicity, establishment of the Radiogenomics Consortium promoted pooling of GWAS data and standardization in reporting GWAS and radiation-induced side effects [14,15]. Now, meta-analysis of aggregate GWAS and whole-genome sequencing continue to promote new discoveries in the field of radiogenomics.
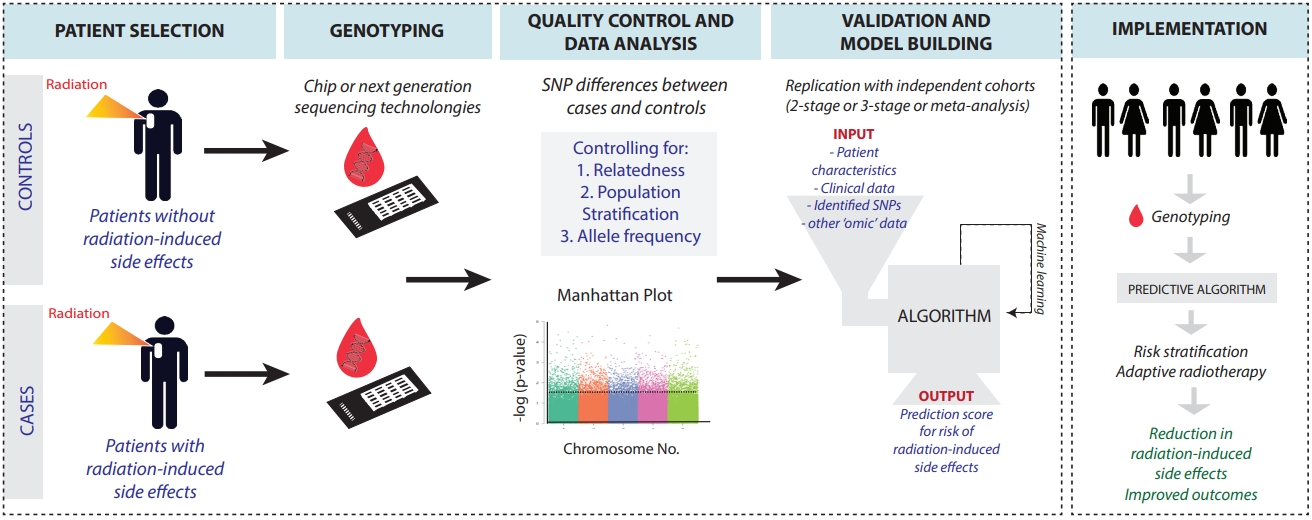
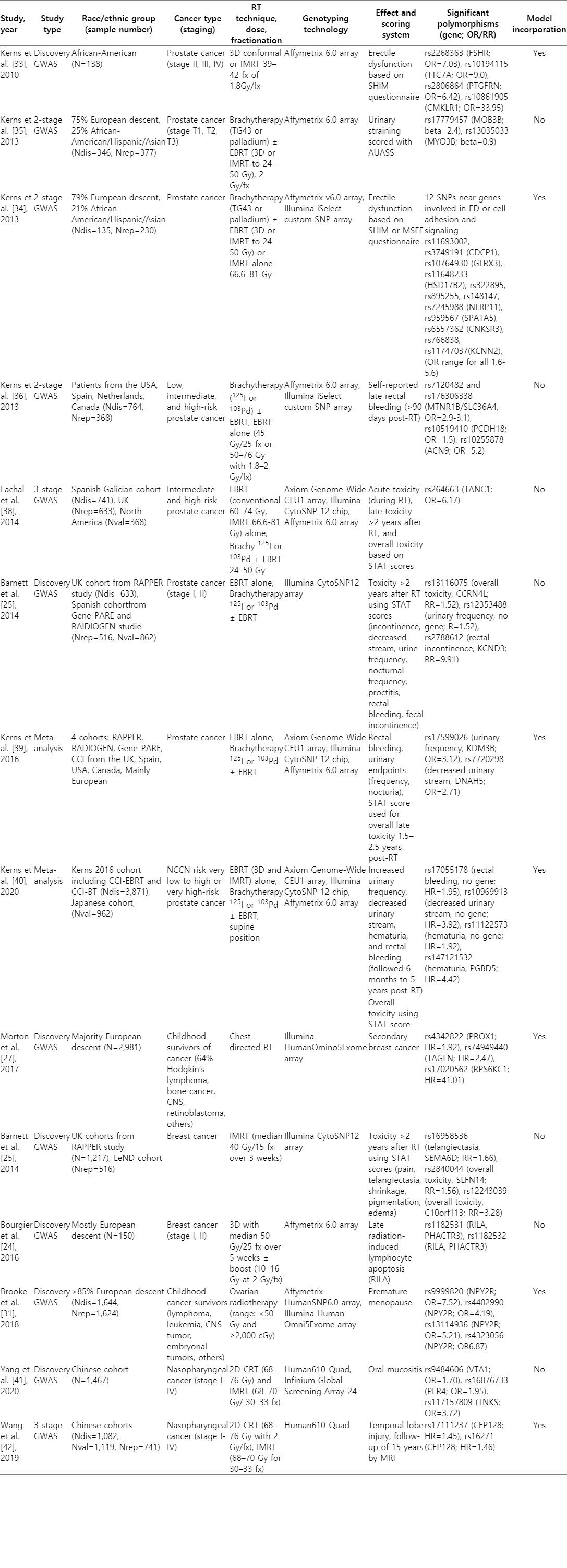
To bookmark the progress of GWAS in elucidating polymorphisms contributing to normal tissue toxicity following radiation, we compiled a narrative review. We focused on seminal literature spanning from 2010 to 2020. Search terms such as polymorphisms, GWAS, radiation, radiotherapy, radiosensitivity, normal tissue toxicity, and radiation effects were queried through PubMed, Google Scholar, and SearchWorks. This review focuses on radiation-induced side effects after chest, pelvic, and head-and-neck irradiation based on data availability. We include both early and late side effects including the risk for secondary cancers. We do not review GWAS identifying polymorphisms affecting development of primary cancers, recurrence, mortality, and tumor radiosensitivity. The GWAS described in this review focus on germline polymorphisms. We hope that early identification of patients that are at high risk for radiation-induced side effects could help to further personalize patients’ treatment plans, including the choice of modality, radiation dose and fractionation schedules. Fig. 1 provides an overview of the GWAS workflow. Table 1 summarizes the GWAS literature presented in this article.
Chest Irradiation
Chest-irradiation puts the breasts, heart, lungs, esophagus, and major blood vessels at risk for toxicity [16,17]. Candidate gene approaches have identified polymorphisms involved in DNA repair pathways related to skin toxicity, pneumonitis, and esophageal toxicity [18–23]. Few studies have used a GWAS approach to discover polymorphisms associated with radiation-induced toxicity. Bourgier et al. [24] attempted to identify polymorphisms influencing late radiation-induced subcutaneous fibrosis, radiation-induced lymphocyte apoptosis, lung toxicity and heart cardiotoxicity using data from the CO-HO-RT trial. Because of the rare event-outcome occurrences and modest heart radiation doses of 2.45 Gy and mean lung dose of 5.3 Gy, they were not able to find any significant SNPs related to lung and heart toxicity. However, they were able to identify SNPs located within the PHACTR3 gene and downstream of GDF-10 that were associated with late radiation-induced lymphocyte apoptosis and radiation-induced skin fibrosis, respectively (Table 1).
To improve the observance of rare events, a composite score that captured multiple radiation-induced side effects was developed. Barnett et al used the scale-independent Standardized Total Average Toxicity (STAT) score as their primary outcome to identify SNPs associated with toxicity two years after radiotherapy in patients that received adjuvant breast radiotherapy and radical prostate radiotherapy [25]. The STAT score included telangiectasia, breast edema, breast shrinkage, pigmentation, and breast pain. GWAS analysis of individual and STAT scores identified rs2881208, rs882460, rs28400440, and rs13116075 SNPs associated with SATB2, CCDC129, SLFN14, and CCRN4L genes, respectively (all risk ratios [RR] >1.5) (Table 1). Though these associations could not validated at a significance level of 0.05—perhaps because of the limited sample size of the validation cohort—a follow-up study by Mbah et al. [26] supported the role of SATB2 rs2881208 in predicting late-radiation toxicity, along with age, supine positioning during radiation treatment, and breast volume.
In addition to broadly identifying genetic risk factors, GWAS have also highlighted differences in genetic susceptibility based on radiation dose. GWAS in survivors of childhood cancers that received ≥10 Gy of chest irradiation identified SNPs near genes involved in cellular proliferation and migration such as PROX1 and TAGLN (hazard ratio [HR] = 1.92; 95% confidence interval [CI], 1.49–2.44) [27]. Whereas, patients that received <10 Gy had different SNP risk factors. Interestingly, the SNPs and associated genes identified in this study did not include SNPs and genes such as BRCA1 and BRCA2 that increase the risk of primary breast cancer. This discrepancy may be due to the smaller sample size of this study compared to general population GWAS. Alternatively, it is possible that the risk factors for breast cancer in the general population may be different from those that are associated with radiation-induced breast cancer. Some studies have shown no association between genes that increase the risk for developing primary breast cancer compared to breast cancer following chest irradiation [12,27]. Large GWAS are therefore needed to identify radiation-specific genetic susceptibility variants and dose-differences.
Genetic susceptibility to radiation toxicity is not only dose dependent but also dependent on when radiation is administered. Studies of survivors of Hodgkin’s lymphoma have shown that the timing of radiation exposure impacts the risk of developing breast cancer and other long-term side effects [28]. If patients receive chest irradiation during puberty when breast development is occurring, they have a higher risk for breast cancer than those patients that receive irradiation post-breast development. Similarly, polymorphisms near the ATG5 and PRDM1 genes were predictors for secondary malignancies for patients irradiated during childhood, but not for those irradiated as adults [29]. Timing of radiotherapy also suggests a role for the circadian rhythm in the development of radiation-induced late side effects. Though identified through a candidate gene approach, circadian rhythm variants, PER3 and NOCT A, were predictive of worse acute and late toxicity if radiotherapy was administered in the morning compared to the afternoon in patients with breast cancer [30]. Normal tissue toxicity is likely fluctuating during a person’s lifetime and may be influenced by baseline health status, such that patients with comorbidities may be at higher risk for developing radiation-induced adverse side effects compared to their healthier counterparts.
Pelvic Irradiation
Radiation and tissue specific genetic susceptibility are also observed in radiation-induced genitourinary toxicity. Female childhood cancer survivors that received pelvic irradiation develop therapy-related premature menopause (PM) at a 10-fold rate higher than their sibling controls. Therefore, efforts have focused on identifying patients that would benefit most from cryopreservation of oocytes prior to starting radiotherapy. Using a two-stage GWAS approach, whereby top-ranked SNPs identified in the first cohort are genotyped in a second cohort, Brooke et al. [31] prospectively followed 799 female survivors of childhood cancer for the development of PM. Adjusting for cyclophosphamide equivalent doses of alkylating agents and ovarian radiation doses, they identified 13 SNPs on Chromosome 4 that increased the risk for PM with an odds ratio (OR) of 25.89 in the discovery phase, and an OR of 3.97 in the replication phase. This haplotype was prevalent in 50% of those with PM, and only observed in 3.8% of controls and in 1.4% of the general female population. SNPs were found near the NPY2R gene, which is known to regulate gonadotropin-releasing hormones pulses, luteinizing hormone levels and ovulation [32]. Variations in expression of genes controlling hormone expression may therefore increase the risk for gonadotoxicity and subsequently lead to premature menopause.
Pelvic irradiation to the prostate in the form of localized brachytherapy, with or without external beam radiation therapy (EBRT), is also associated with radiation-induced side effects in 5%–10% of patients. Side effects can include erectile dysfunction (ED), rectal bleeding, urinary and fecal incontinence, decreased urinary stream, and proctitis. The first GWAS looking at radiation-induced side effects in prostate cancer studied ED risk [33]. GWAS identified SNPs located near genes involved in cell adhesion and cell matrix association, rather than radiation-affected pathways, such as DNA damage repair [34]. There were also group enriched SNPs near the FSHR gene that increased the risk for radiation-induced ED in African Americans [33]. This study and others revealed that increasing the number of alleles increases the risk of erectile dysfunction [33,34]. Younger men with more risk alleles had a similar likelihood of developing ED as older men with fewer risk alleles. One allele had an OR of 2.2, while multiple alleles had an OR >2.2 [34]. Furthermore, incorporation of SNPs into a multivariate model along with ancestry was more accurate in predicting ED than clinical factors alone such as stage, Gleason score, and pre-treatment sexual function (area under the curve [AUC] for genetic model = 0.983 vs. AUC for clinical model = 0.749) [33]. Though not directly addressed in this study, incorporation of both genetic risk factors and clinical metrics can increase the predictability of these models.
GWAS have also been used to identify polymorphisms associated with changes in late-toxicity such as changes urinary function, rectal bleeding, and overall toxicity. Eight SNPs, including one associated with MYO3B (rs13035033), increased the risk for urinary straining [35]; though this finding could not be replicated [37]. GWAS of individual endpoints such as rectal bleeding identified SNPs near genes regulating vascular proliferation [36], while markers of overall toxicity identified TANC1, a gene involved in muscle regeneration, as a risk factor [38]. To increase the power to detect rare polymorphisms, Kerns et al. [39] used a meta-analysis approach to identify SNPs associated with increased urinary frequency, rectal bleeding, decreased stream and overall toxicity. This study validated previous SNPs, discovered new SNPs, and most notably it showed that multiple data sets could be combined to increase the power of GWAS. A follow-up meta-analysis study included additional samples from a Japanese cohort to identify new polymorphisms associated with hematuria, and validated SNPs previously associated with increased urinary frequency, decreased urinary stream, and overall toxicity [40]; again emphasizing the power of analyzing GWAS datasets in aggregate.
Head and Neck Irradiation
Oral mucositis is an acute complication of head and neck irradiation. To identify patients that would be at risk for adverse oral mucositis outcomes, Yang et al. [41] performed a GWAS in more than 1,000 nasopharyngeal carcinoma patients receiving radiotherapy. Using gene set analyses of their identified variants they were able to link telomere biological processes to radiotherapy toxicity and thereby illustrate the use of GWAS in enhancing our understanding of radiobiology.
Temporal lobe injury is a late radiation-induced side effect in patients with nasopharyngeal carcinoma. Genetic susceptibility studies for radiation-induced central nervous system toxicity are starting to emerge. Wang et al. [42] conducted a three-stage GWAS study with more than 1,000 patients for each stage. Their analysis identified SNPs near the gene CEP128, which codes a centrosome protein involved in cell cycle progression, and is associated with temporal lobe injury risk (HR=1.45; 95% CI, 1.26–1.66). Furthermore, they discovered that 12.7% of the GWAS-based genes were associated with the neurogenesis pathway. SNP (missing period) differences associated with the CEP128 gene were also associated with changes in levels of expression [43]. These results suggest again that radiation-induced injury is likely affecting tissue specific cellular pathways.
Models of Radiation Toxicity
GWAS have provided novel insight into radiobiological processes not otherwise recognized through candidate-gene approaches. They have also highlighted the differences in genetic susceptibility due to radiation dosing and timing of administering radiation. GWAS have distinguished polymorphisms comparing tissue-specific toxicity versus overall toxicity and acute versus late radiotherapy side effects. Two proposed models may in part explain the complexity of genetic susceptibility to radiation—a deterministic model and a stochastic model [10,44]. Under a deterministic model, each tissue can have its own threshold for toxicity and these thresholds can vary between patients. In a stochastic model, radiation may impact cellular function through general processes such as increasing reactive-oxygen species and promoting genomic instability. These processes would nonspecifically affect pathways that increase the risk of radiation-induced side effects. Variability in observing deterministic or stochastic processes would depend on individual patient genetic vulnerability. Patients with polymorphisms that increase stochastic events may be at higher risk for overall toxicity, whereas patients with polymorphisms affecting tissue-specific pathways would manifest more restricted side effects. Phenotypic expression of adverse effects would thus be dependent on genetic susceptibility and life style factors. For example, if the patient is already at risk for developing erectile dysfunction, both genetically and because of lifestyle factors, exposure to radiation may be the last “hit” required for the development of ED through a deterministic or stochastic process. GWAS and candidate-gene approaches have been successful in identifying “general” toxicity polymorphisms and side-effect specific polymorphisms. Integration of cellular assays, “omic” data such as transcriptome-wide association studies (TWAS) and proteomics will aid in distinguishing between the two processes at the patient level.
GWAS Challenges and Opportunities
Despite recent progress and its promise to personalize patient care, integration of GWAS findings into clinical settings remains a challenge. Reproducibility is often hard to achieve because of small sample sizes, which decrease the statistical power needed to detect moderate-to-weakly associated polymorphisms, and high false positives from multiple-hypothesis testing. To address these challenges, the Radiogenomics Consortium encouraged standardization of reporting of GWAS, central pooling of large databases, and standardization in reporting acute and late side effects [14]. Combining multiple datasets, however, posed additional statistical challenges that have now spawned new mathematical models and statistical approaches. These mathematical models have been applied with success to predict radiotherapy complications such as late rectal bleeding and erectile dysfunction and have identified biological processes of radiation damage [45,46].
Incorporation of GWAS findings into machine learning also has the potential to increase the clinical utility of GWAS. Lee et al. [47] used machine learning-based multivariate modeling to predict four urinary symptoms following radiation therapy. They found that weak stream was predicted when the top 75% of SNPs were included in their model compared to a model that only included clinical predictors (R2 of 0.80 vs. R2 of 0.60). Machine learning algorithms can be applied to integrate genetic data, clinical correlates, and patient characteristics to build models that can evolve as patients’ characteristics change through the course of treatment [48]. However, in order for models to be truly predictive, racially heterogeneous populations must be included.
It is known that polymorphisms can vary by race/ethnic groups. For example, SNPs associated with the TGFb1 promoter and the NFE2L2 promoters were associated with late radiation effects in African Americans, but not Whites [49]. The failure to include diverse patient groups in GWAS has two main effects: (1) decreased discovery potential and generalizability and (2) decreased predictive power of model building. Although there is concern that inclusion of non-European populations increases the signal-to-noise ratio, low frequency variants or variants that are completely absent in European populations may be missed [50]. In addition, the effect sizes of identified variants in one group may be the different in other populations. Wojcik et al. [51] used a large data set that included Hispanic populations, African-American, Asian, Native Hawaiian, Native American, and multi-ethnic individuals to show that the effect size of published trait-variant associations were significantly weaker in Hispanic and African-American populations. Predictive models based on these effect sizes and variants could potentially result in erroneous predictions that could further increase already existing health disparities. Thus, recruitment of under-represented groups, development of algorithms robust to the inclusion of heterogeneous populations, building of databases, and integration with other “omic” data, will strengthen the clinical potential of GWAS.
Conclusion
Since its adoption in radiation oncology, GWAS have identified polymorphisms that increase genetic susceptibility to radiation-induced tissue toxicity. However, challenges in high false rate discoveries due to multiple hypothesis testing and small sample sizes have failed to reproduce some results. Development of techniques to score side effects and standardize GWAS reporting, along with the establishment of the Radiogenomics Consortium have partly overcome some of the challenges inherent in GWAS. Meta-analyses of GWAS now include larger patient cohorts, more diverse populations, multiple steps of validation, and new statistical approaches. Incorporation of genetic susceptibility of normal tissue, tumor radiosensitivity, clinical markers, and patient characteristics into adaptive models is poised to make genomically-guided radiotherapy a reality.
Notes
Conflict of Interest
No potential conflict of interest relevant to this article was reported.